cBioPortal is an interactive open-source platform that allows users to visualize and analyze genetic data from cancer studies. The platform includes the analysis of DNA mutations, gene expression, fusion genes, and copy number alterations, which can be viewed in context of clinical parameters such as sex, age, smoker/non-smoker etc.
CUBI generally delivers inhouse cancer variant analysis results through cBioPortal, however externally generated analysis can also be imported into our local instance.
Resource
You can find the internal cbioprtal instance at cbioportal.bihealth.org. Please note this is only accessible from internal networks, and a valid Charité or MDC account is required to log in. We support both public and private projects. Access to the private projects needs to be approved by the PI in charge of the data.
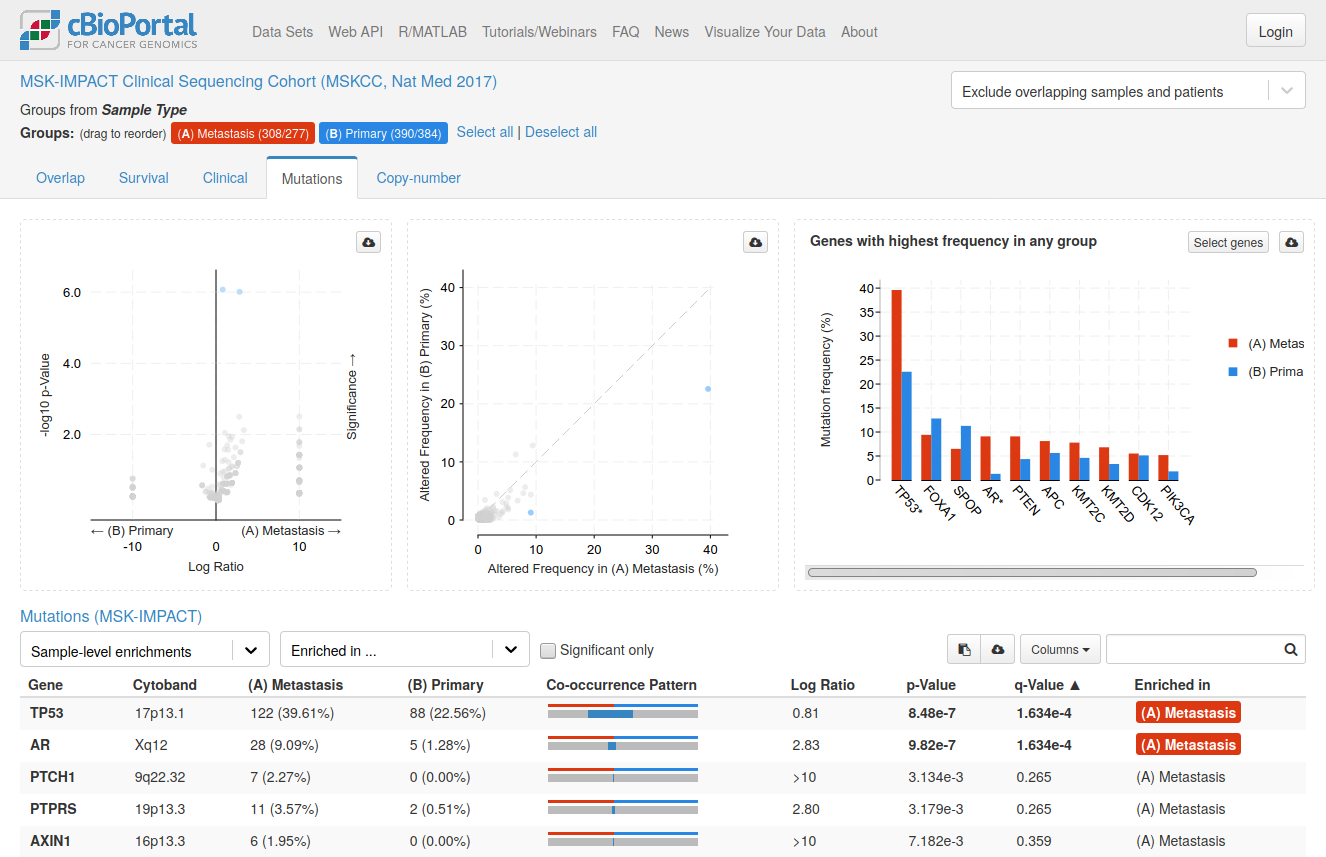
cBioPortal showing the comparison of primary and metastatic prostate cancer on a public data set.
Services
Private Study Import
Do you have tumor data (either with or without clinical information) that you
want to analyze in cBioPortal? Access to individual studies are given
in an orthogonal way. Studies can be part of a lab with one
responsible PI. All registered members have access to all studies of
this lab. For collaborations, studies can also be placed within
projects, where access can be granted independently of lab membership.
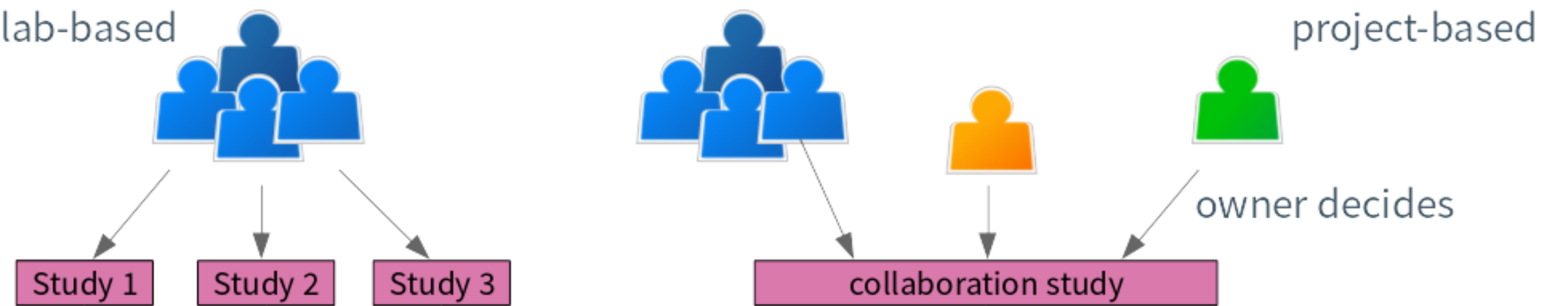
Access to studies in cBioPortal is based on labs or projects.
Contact us to learn how you can get the data into the in-house cBioPortal instance.
Public Data Import
Are you interested in seeing your studies in the context of additional publically
available studies? We are happy to import one of the 100+ public
studies into our instance so you
can do this!
Consistent Upgrade Cycles. cBioPortal is continuously developed and maintained by a multi-institutional team (http://www.cbioportal.org/about) with frequent updates. In order to provide the most recent cBioPortal features we are regularly running a two-step upgrade of the local instance.
Workshops
We are happy to organize workshops for explorative data analysis using
cBioPortal. Please also refer to the following webinars at
http://www.cbioportal.org/tutorials .
Contact
Please contact cubi-helpdesk@bih-charite.de for more information on how to obtain access and use this resource.
Last modified: Feb 7, 2023